Medicine can be as much art as science, a detective story in which doctors rely not only on lab tests and x-rays, but on their own experience and clues from a patient’s history to develop diagnoses or predict future health problems. But all of those lab tests, blood pressure readings, magnetic resonance imaging (MRI) scans, electrocardiograms, and billing codes add up to reams of data, which before too long will be joined by individual gene sequencing. Computer scientists are increasingly applying machine learning techniques to all that data, searching for patterns that can aid diagnosis and improve clinical care.
“Machine learning plays, I think, an essential role in medical image analysis nowadays,” says Kenji Suzuki, assistant professor of radiology and medical physics at the University of Chicago’s Comprehensive Cancer Center. Suzuki has been working on automating the detection of cancerous lesions in images from x-rays or computed tomography scans. Considering that radiologists may miss 12%30% of lung cancers in such scans, a machine learning tool offers great potential.
Since the mid-1980s, computer scientists have tried to improve on that performance using feature-based machine learning in which the computer would pick out morphological features, texture differences, and more to identify abnormal tissue. But such cataloging of features still misses some cancers that doctors are able to spot with their own eyes. Sometimes the features the computer is seeking can be subtle or overlap with normal anatomical structures such as bone, making them more difficult to spot. So Suzuki asks the machine to instead focus on the values, such as intensity, of individual pixels. “Because the computing power has increased dramatically in recent years, we can process the pixel values directly,” he says. The resulting system is highly sensitive, achieving up to 97% accuracy.
As the use of electronic medical records gains acceptance, machine learning is likely to play an even larger role in clinical medicine.
But one concern is making the program so sensitive that it starts finding nonexistent lesions. In Suzuki’s lung cancer tests, the feature-based algorithm falsely identified five lesions per patient, while the pixel-based method produced less than one false positive per patient and no false negatives. Suzuki says each method produces a different type of false positive, so combining the two approaches leads to the most accurate outcome. Suzuki is now working on expanding the technique to other types of cancer and other imaging methods, such as MRI and PET. “You just need to train the machine learning technique with new images,” he says.
A Guide for Diagnosis
At IBM Almaden Research Center, the Advanced Analytics for Information Management (AALIM) project applies machine learning to a wide variety of data—readings of vital signs, tests such as echocardiograms, and demographic information—to chart the medical histories of patients over several years. By comparing the history of hundreds or thousands of people, the system can identify previous patients who are similar to a current patient, then apply collaborative filtering to suggest the best diagnosis and treatment options for the new patient.
“With a large number of pre-diagnosed patient datasets available in electronic health records, physicians can now benefit from the opinion of their peers on cases similar to their patients,” says Tanveer Syeda-Mahmood, head of IBM’s Multimodal Mining for Healthcare project. The hope is that by helping doctors base their decisions on quantitative information, the number of diagnostic errors can be reduced. AALIM, Syeda-Mahmood says, provides “a holistic view of the patient’s condition,” producing one-page summaries, long-term profiles of various measurements of health, and detailed comparisons showing diagnosis, treatment, and outcomes for similar patients.
The computer might, for instance, help a relatively new doctor decide that she needs to consult a specialist. Approximately 5% of cases, says Syeda-Mahmood, are ambiguous enough that even senior clinicians ask other doctors for their opinions, which AALIM easily provides. In emergency rooms, it can cut the time doctors spend flipping through charts by half. Although researchers have tested the system on patient data, it would likely require approval from the Food and Drug Administration before it could be used in a hospital.
Finding ways to use all this medical data often requires new developments in machine learning. For instance, Syeda-Mahmood wanted to give the computer a doctor’s ability to recognize some types of heart disease by the characteristic shape of waves produced by an electrocardiogram. She developed a new function, called a constrained non-rigid translation transform, which could identify the similarity between shapes in different ECG readouts.
The Power of Prediction
Although diagnosis and treatment are key aspects of medical care, prediction is also important, especially when it can lead to early interventions. We all know, for instance, that factors such as weight and blood pressure can give an idea of a person’s risk of heart disease. But those sorts of risk scores are based on population-wide models, says Shyam Visweswaran, assistant professor of biomedical informatics at the University of Pittsburgh. “If you build a model from a group of people who are kind of similar to the current patient, you might do better,” he says. Visweswaran has developed an algorithm that lets a computer use clinical data to learn a model tailored to one specific patient and predict outcomes for that person.
The computer takes all the data it has on the patient, such as age, blood pressure, and lab results, and then picks one variable and builds a model of all the patients in its database who share that variable. It could, for instance, compare everyone in the 5055 age group. It builds a model for each variable it can find, looks at which ones best fit the patient at hand, and then averages the best models to make a personalized prediction of that patient’s outcome. Whereas a population model only uses a handful of variables considered to be the best—it could be a simple checklist of several risk factors, for instance—this approach can potentially use any of hundreds of variables. One additional advantage is the machine might identify some factor that is predictive, but that medical science was not previously aware of, opening up new areas for research, Visweswaran says.
“With a large number of pre-diagnosed patient datasets available in electronic health records, physicians can now benefit from the opinion of their peers on cases similar to their patients,” says Tanveer Syeda-Mahmood.
As with Suzuki’s pixel-based processing, this is another machine learning method that has benefited from the growth of processing power. As recently as five years ago it might have taken a half-hour to build all these models. Today it takes less than a minute, so the machine can guide diagnoses in real time during patient visits.
This approach could help predict an intensive-care patient’s risk of an infection spreading to other organs, which is a notoriously difficult task, and lead to earlier or more aggressive treatment. It might help doctors decide, for instance, which pneumonia patients need to be admitted to the hospital and which patients could be sent home with antibiotics.
In the informatics program at Children’s Hospital Boston, assistant professor Ben Reis and his colleagues are working on predicting a patient’s future diagnoses years in advance. They have developed Bayesian models that they call Intelligent Histories, which comb through the standard diagnostic codes used for billing, to find patterns in a patient’s history that predict risk. In their first application of the work, they discovered they could identify patients at risk of domestic abuse as much as two years before the doctors seeing those patients first discovered the problem.
Doctors are supposed to screen patients for domestic abuse, but often miss it until the problem becomes acute, says Reis. Not only can the computer aid in screening for known signs of abuse, it also picked up other diagnostic codes in the test that had not been thought of as predictive, such as infections, which might teach doctors something about domestic abuse.
The Children’s Hospital Boston team is working to expand their modeling to other types of diagnoses. At the same time, they are refining the machine learning itself. For instance, “we’re trying to quantify how the quality of the data that goes into the model affects the results that come out,” says Reis.
As the world moves to a greater use of electronic medical records, machine learning is likely to play an even larger role in clinical medicine, researchers predict. Visweswaran says genetic data, in particular, is going to require complicated computational models if it is going to be of value. Soon, experts expect, the cost of gene sequencing will drop to the point that individual genomes will become part of people’s medical records, and will be available to the same data mining and pattern recognition approaches being applied to other data.
Genetic data will be too complicated and too voluminous to be handled with old-fashioned charting systems. “You have to have computational tools to query this data,” Visweswaran says. “There’s no way it can be done on paper.”
Gruhl, D., et al.
Aalim: A cardiac clinical decision support system powered by advanced multi-modal analytics, International Medical Informatics Conference, Cape Town, South Africa, Sept. 1215, 2010.
Reis, B.Y., Kohane, I.S., and Mandl, K.D.
Longitudinal histories as predictors of future diagnoses of domestic abuse: modeling study, British Medical Journal, 339, Sept. 2009.
Syeda-Mahood, T., Beymer, D., and Wang, F.
Shape-based matching of ECG recordings, 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, August 2327, 2007, Lyon, France.
Suzuki K., Zhang J., and Xu, J.
Massive-training artificial neural network coupled with Laplacian-eigen function-based dimensionality reduction for computer-aided detection of polyps in CT colonography, IEEE Transactions on Medical Imaging 29, 11, Nov. 2010.
Visweswaran, S., Angus, D.C., Hsieh, M., Weissfeld, L., Yealy, D., and Cooper, G.F.
Learning patient-specific predictive models from clinical data, Journal of Biomedical Informatics 43, 5, Oct. 2010.
Figures
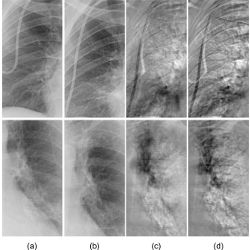
 Figure. Kenji Suzuki and colleagues’ comparison of their rib-suppressed temporal-subtraction (TS) images with conventional TS images: (a) previous chest radiographs, (b) current chest radiographs of the same patient, (c) rib-suppressed TS images with fewer rib artifacts, and (d) conventional TS images.
Figure. Kenji Suzuki and colleagues’ comparison of their rib-suppressed temporal-subtraction (TS) images with conventional TS images: (a) previous chest radiographs, (b) current chest radiographs of the same patient, (c) rib-suppressed TS images with fewer rib artifacts, and (d) conventional TS images.




Join the Discussion (0)
Become a Member or Sign In to Post a Comment