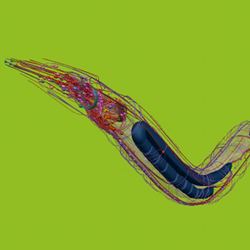
Imagine if biology experiments that once took months to set up and run in a laboratory could be reliably duplicated in minutes with a few clicks of a computer mouse. That is the goal of a diverse international team of computer scientists and neuroscientists in the OpenWorm Project, who are developing a comprehensive in silico model of C. elegans, a tiny, soil-dwelling nematode.
In addition to shedding light on how various biological systems—neural networks, muscles, sensory organs and so on–work and interact, they hope that OpenWorm will provide a platform for applications such as new drug discovery and disease research.
As creatures go, C. elegans is simple, with just 959 cells and 302 neurons. Yet when one considers combinations of these things, and the internal workings of cells, the number of possible behaviors and interactions becomes astronomical. Indeed, no complete molecular model of any one cell has ever been constructed. "We are dealing with a very complex system," says project coordinator Stephen Larson, a computational neuroscience consultant.
OpenWorm will be a tightly integrated set of four or five models of the worm’s major biological systems. The first of these, to be rolled out in about a year, will use two algorithms to model the animal’s neurons, their electrical interactions with the muscles, and the muscles’ response. A good model will predict with reasonable accuracy how a live worm crawls under various conditions, Larson says.
The Base Model
Larson says that this first model will be "fairly coarse-grained." Follow-on work will refine the neuro-muscular model to consider sub-cellular components such as proteins and DNA. Indeed, he says, one of the most daunting design challenges of the overall effort is deciding at what levels in biology the models must work in order to produce reasonably accurate results. He says the accuracy of models in terms of their ability to describe the inner components of live organisms, and their ability to predict behavior, are orthogonal characteristics.
"One of the things the project is trying to do is answer the question of how little biological realism do you have to have to explain a lot of behavioral data."
Subsequent OpenWorm modules–one for the worm’s sensory organs, for example–will follow. Development and integration of these components is being enabled by a development framework called Geppetto (after the fictional woodcarver who made the puppet Pinocchio), which is emerging under the direction of OpenWorm team member Matteo Cantarelli, a software engineer and research associate at University College London.
"Geppetto will be a platform that is generic enough that it can be driven by different data and models," Cantarelli says. "We are using OpenWorm and C. elegans as a case study for what kinds of different modules have to exist and what the interactions between them will be."
One of the tricks that Geppetto must perform is integrating and harmonizing the vastly different spatial and time scales that components will operate at, as well as dozens of different software packages and programming languages. It must be algorithm-agnostic, assuming different people, some external to the project, will code components. A key objective of Geppetto, indeed of all of the project’s work, is that it be completely open source, Cantarelli says. "The idea is for it to be a real engineering product that can be reused and improved by others."
The OpenWorm core team consists of about 10 people in the U.S., Russia, Great Britain, and Ireland using a variety of sharing tools, such as Google Hangouts and Dropbox. "All the work we do is instantly available online, and all our meetings are streamed live on the Internet," says Mike Vella, a doctoral candidate in neuroscience at the University of Cambridge. "The huge benefit is people coming along and saying ‘I can help,’ and then submitting some code or an idea."
Modeling Science
To make a good biological model requires a good knowledge of the underlying science, Vella says, with gaps or errors in the science resulting in corresponding gaps or errors in the OpenWorm models. However, he adds, "If there are significant unknowns in the science, our model will still hopefully act as a motivator for other biologists and modelers to fill in those gaps. The advantage of a good model is that it provides boundaries on what we know."
The genetic structure of various mutants of C. elegans is known well, and that will be a springboard for further modeling with OpenWorm, Larson says. "These known gene deletions give you some insight into diseases. So you can go from models that make predictions about healthy cells to models that can predict diseases or dysfunctions."
Modeling even a relatively simple organism like the 959-cell worm won’t be a job for a desktop PC. For example, a custom-designed fluid mechanics simulator that works at the particle level performs linearly with the number of processors devoted to it, Vella says. Developers are now using workstations fitted with multiple NVIDIA Tesla graphics processing units (GPU), but "in the future, we are going to have to think about supercomputers," he says.
Indeed, computer hardware will be a big issue in any plan to follow up the C. elegans model with models of, say, a mouse heart or a fruit fly. "The computing resources that we need to do the worm are so great that we are not sure if adding 100 times more cells is feasible," Larson says. His team is exploring the use of cloud computing for processes that are highly parallelizable, and supercomputers for processes that are computationally intensive. "We are looking at a fairly heterogeneous computing framework, some parts of which optimize memory, some compute time, and some interconnect," he says.
Gary Anthes is a technology writer and editor based in Arlington, VA.



Join the Discussion (0)
Become a Member or Sign In to Post a Comment